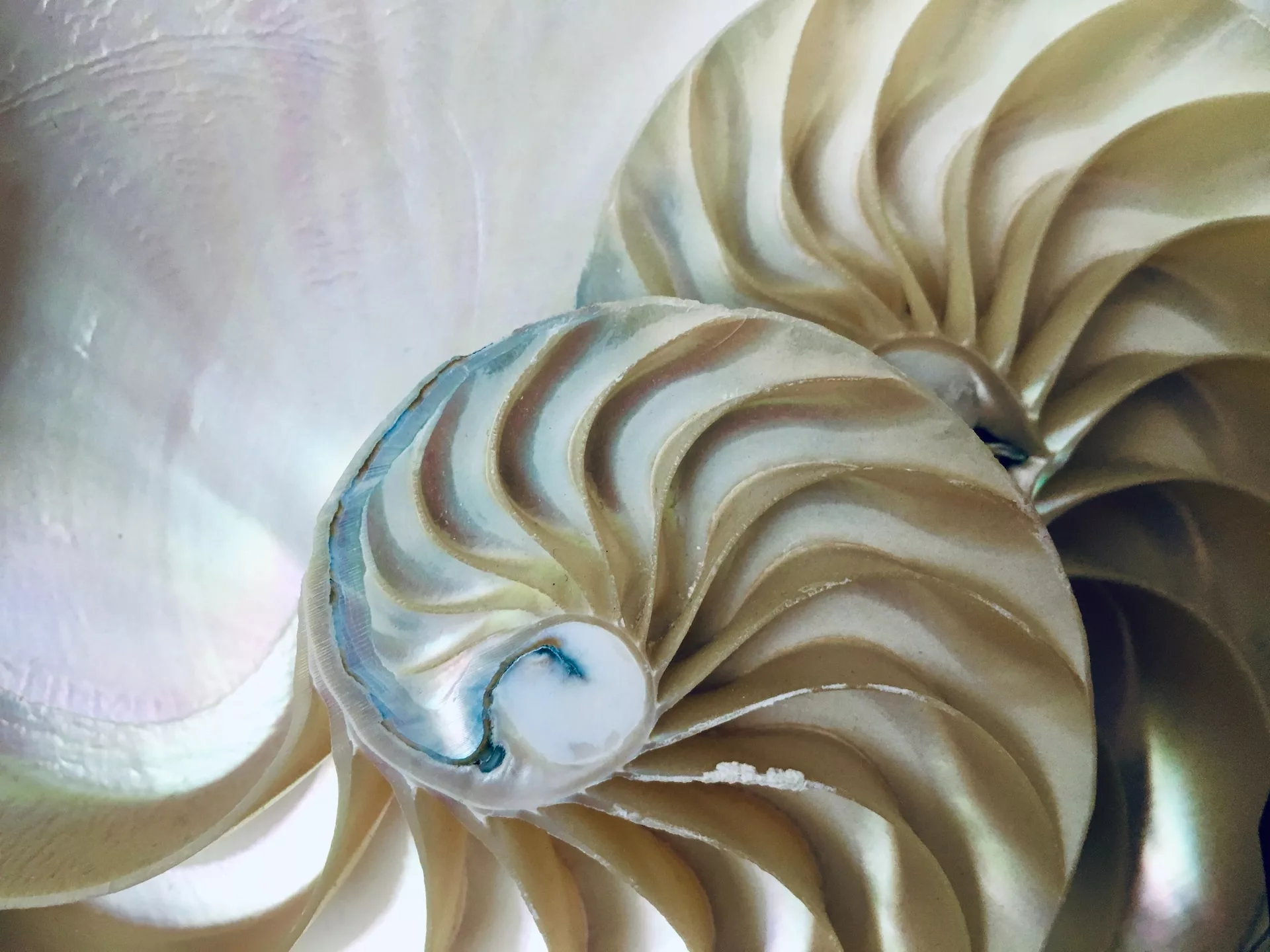
Biology and mathematics are closely intertwined. Classically, the fibonacci sequence was used to explain the development of many species. Now, AI is necessary to understand how this development arises. Photo credit: Giulia May via Unsplash
One of the earliest mathematic models which found expression in biological systems is the Fibonacci sequence. The sequence is quite simple: it consists of a series of numbers where each new number in the series results from the addition of the previous two. An interesting feature of this sequence is that the ratio of adjacent values approaches a limiting value, given the symbol τ, which is often referred to as the “golden ratio”. The name came about because the golden ratio is a visually appealing proportion and is, therefore, consistently found in ancient art and Greek architecture. Surprisingly, it also occurs in biological contexts: sunflower seeds form a progression of helices that encode this ratio. Each seed is placed at a rotation angle of τ from the previous one. It is thought that Kepler was the first to realise this.
Since Kepler’s time, developmental biology—the field of biology concerned with describing how plants and animals grow—has evolved enormously. Although it began with noticing the prevalence of geometric patterns in living things, the question has since turned to one of modelling: scientists are interested in how the emergent properties of a system, such as the specific pattern of sunflower seeds, arise from the interaction of individual parts—proteins and other biomolecules. What makes it even more interesting, and challenging, is that the composition and structure of these thousands of molecules within each of the approximately 1014 cells in a human body are all entirely specified by the DNA code.
Modelling, together with computer simulation, hopes to fully understand the process of developing a functioning organism given only the DNA sequence. When an organism is developing, almost identical collections of cells form spatial patterns without any external stimulation, which result in features that are characteristic of that organism, such as the stripes on a zebra’s coat. Understanding how the cells “know” to organise themselves in this manner involves modelling processes within a single cell, as well as the interactions between cells.
Abstracting away that detail [of biological processes], however, leaves us with predictions that are too rough compared to experimental data.
Doing so, however, presents a significant challenge because of the scale of simulations that are involved: many of the processes that we aim to model contain too much detail for computers to handle. Abstracting away that detail, however, leaves us with predictions that are too rough compared to experimental data. This is where the field of AI, or artificial intelligence, comes in
AI and the subsidiary field of machine learning, abbreviated to ML, have proven useful in tackling highly complex problems on which more traditional techniques are not able to do particularly well. The success is largely down to the manner in which an ML algorithm works. Broadly speaking, ML is an approach in which, rather than coding an algorithm to solve a particular problem, the machine or computer is provided with data and tasked with coming up with its own algorithm. This contrasts the traditional techniques, in which the scientist hard-codes the rules themselves. These “rules” are often inspired from real processes which occur in nature and are meant to mimic these processes, a method of modelling known as mechanistic modeling.
Part of the success of ML in recent years is due to a particular type of learning technique called Deep learning. Here, a large amount of data is fed into the neural network to “train” the network to make predictions or decisions when faced with new data. Not too surprisingly, research has started to look at developmental biology as a potential field of application. Although the predictive power that we gain from doing so is potentially huge, the downside is that these methods do not necessarily help us gain an understanding of the underlying mechanism at hand or how the real-lifesituation unfolds. Although this seems to limit the usefulness of such methods to examples where the underlying mechanism is already known, this is not necessarily true.
Modelling processes within cells: Protein folding
Predicting the three-dimensional shape of a protein from only the order in which its molecules are bonded together is called the “protein folding problem”, and progress in the area is assessed every two years by the CASP experiment. A protein is formed by the bonding together of many molecules called amino acids. These molecules have a central carbon atom and various side chains. It is these side chains that determine how a particular protein spontaneously folds into the three-dimensional shape that it will conform to for the rest of its lifetime. This shape is crucial for determining which molecules the protein will interact with; knowing the shape not only determines its function but assists researchers in designing drugs that bind to proteins based on this shape. Ideally, creating a model of this process would require detail at the atomic level so that folding would result from a detailed force field. Including this amount of detail, however, is infeasible, as the rules of folding are too complex for any computer—including today’s supercomputers—to compute.
To overcome this limitation, DeepMind developed a deep learning algorithm called AlphaFold, which predicts a protein structure given only the amino acid sequence. In the 14th iteration of CASP, known as CASP14, AlphaFold significantly outperformed other predictions, with DeepMind observing that ‘it regularly achieves accuracy competitive with experiment’. This is an extraordinary feat, considering the complexity at hand. However, how the algorithm reaches the final folded protein shape—the particularities of the folding process—is not given to us as an output of the algorithm.
In previous methods for predicting protein shape, humans developed the simulation rules. As such, there was no obscurity in the process by which these algorithms achieved their final answers. For example, Lattice models are simplified, or reduced, models that consider the properties of amino acid side chains as a whole rather than the properties of the atoms themselves, thereby simplifying the problem by reducing the number of interactions that need to be considered. An obvious drawback is that these methods cannot (even nearly) reach the accuracy of AlphaFold.
In this example, however, knowing how the simulation arrived at the final answer is not so important. This is because we do know how it is done in practice (or in nature): it is the electrostatic interactions of the atoms on each amino acid that causes the protein to fold into the shape, which minimises the repulsion between atoms.
There are, however, cases where we do not know how a certain feature comes about, particularly in fields where properties are emergent. An emergent property is the result of complex interactions between many tiny components to produce behaviours or properties which the individual components do not themselves possess. It is in these cases where it makes less sense to replace current methods with machine learning methods. This is because, while the deep learning model could predict the emergence of certain properties, and perhaps to high accuracy, this does not provide an answer as to how the property emerged—doing so requires finding the underlying mechanism. This process of proposing a particular mathematical model that can be experimentally verified or falsified is known as mechanistic modelling.
Modelling whole cells: Morphogenesis
Morphogenesis is the development of the shape and form of an organism from almost homogenous tissue. Homogenous tissue consists of cells that are entirely the same and not yet specialised to fill a particular function. D’Arcy Thompson said that ‘the organism is the τελoς, or final cause, of its own processes of generation and development’. An organism seems to “know” what shape to become, and yet the instructions for the process are not contained in the DNA code. Morphogenesis is a complex process but, importantly, is the result of the interaction of various molecules and cells.
In 1952, Alan Turing, an English mathematician, wrote a paper on “chemical morphogenesis” in which he showed that the development of patterns in tissue can be predicted using reaction-diffusion models. Reaction-diffusion models are a type of mathematical model in which cells either attract or repel those around them until the configuration that causes the least repulsion is achieved. For example, reaction-diffusion mechanisms have been used in studying animal coat markings to show that the shape of an animal’s tail determines what markings it carries. In particular, reducing the cross-sectional diameter down the tail ‘forces the spots to become stripes’, such as in the spotted cheetah. The model predicts that the opposite cannot be true—that stripes cannot turn into spots down the length of the tail— and no animal in nature is yet to be observed with such a pattern.
More recently, with increased computational power, computer models called cellular automata have started to be employed in modelling more complex morphogenesis mechanisms. A cellular automation is a kind of decentralised computer, which contrasts with our usual idea of modern computers in which there is some central processing unit sending out instructions. The cellular automata are composed of several units, called cells, placed on a grid structure. Each cell can then be in one of many states. There are certain rules as to how cells update according to the state of their neighbours, which are called “update rules”. Importantly, these contrast machine learning methods because the rules of the game are deterministic and known by us.
…computer models called cellular automata have started to be employed in modelling more complex morphogenesis mechanisms.
One of the most famous cellular automata is John Conway’s Game of Life. Each cell in an infinite two-dimensional grid (in practice, this is a large grid) is instantiated to one of two states: either “alive” or “dead”. To visualise the automaton, we represent an empty cell as dead and a filled-in cell as alive. Each cell is then updated according to one of four rules, depending on the states of its eight neighbours (above, below, beside, and on the diagonals). For example, an alive cell with fewer than two alive neighbours transitions into the dead state. These rules are applied to every cell in what is called a time step, producing a new grid of alive and dead cells. This process is repeated for many time steps, eventually resulting in different configurations of black and white cells on the grid. Still life configurations are those that result in a stable configuration of cells that does not change in time, even as rules are applied again and again. We can get other types of stable configurations, where the grid oscillates between several different states or even moving configurations, which have been coined ‘spaceships’, because the filled-in cells propagate across the grid.

The key point about cellular automata is that the computation is decentralised, in the sense that decisions are made based on the local arrangement of specific cells, rather than via some central processing unit which sends out instructions. This is precisely the fashion in which morphogenesis proceeds: chemical units called morphogens occur all over our body and are responsible for sending out signals to cells to control cell development.
In a 2020 study, researchers trained a neural network to “learn” the update rules of a cellular automation, which can mimic organism development. Here, cell states are vectors of 16 real values (rather than a binary choice of black or white, as in the previous example), representing the colour and other parameters of the organism. Perturbations to the final organism result in the regeneration of that part of the image. The algorithm works by at first allowing all update rules to occur with equal probability. The result, however, is that the original image will not be regenerated. In an intelligent way, the machine learning algorithm can deduce, after many rounds, the weights of each update rule—or the probabilities with which each occurs— so that the cellular automata, when run, reproduces the original image or organism.
To put things into perspective, the neural network that is used to learn the update rule has approximately 8,000 parameters. Thus, the number of possible update rules is extensive. The idea of using statistical methods to learn the update rules of a cellular automata, as opposed to “hard-coding” them from the beginning, is not a new one and is already prevalent under the name of genetic algorithms. It finds application in, for example, writing computer programs for tasks, such as coding a robot to pick up litter in the most efficient way. This is the first time, however, that researchers have managed to utilise it in developmental biology.
On the one hand, it seems that there is a lot of potential for the use of AI, and specifically machine learning, in developmental biology. On the other hand, the complexity of the field highlights the shortcomings of machine learning methods. While they are useful for training certain aspects of the model, if it is the underlying mechanism that we wish to understand, we often must instead turn to mechanistic modelling. The parametrisation of mechanistic modelling, however, such as that discussed above in cellular automata, offers a kind of middle road between the two, in which mechanistic models can be parameterised and therefore tested with the help of machine learning tools.